Machine learning extracts marks of thiamine’s role in cold acclimation in the transcriptome of Vitis vinifera.
written by Konecny, T.; Nikoghosyan, M.; Binder, H.,
- 1Armenian Bioinformatics Institute, Yerevan, Armenia
- 2Interdisciplinary Centre for Bioinformatics, University of Leipzig, Leipzig, Germany
- 3Bioinformatics Group, Institute of Molecular Biology Institute of National Academy of Sciences RA, Yerevan, Armenia
INTRODUCTION
Vitis vinifera is a species of grapevine plant that has been cultivated for thousands of years. Its fruit is used both in the production of wine and as a table grape. It is one of the most economically important fruit crops globally. Of the thousands of varieties of grapes, only a few are of commercial significance for wine and table grape production. The European grapevine is adapted to various climatic conditions, but its performance is best in temperate regions with mild winters and warm summers. The greatest challenge that climate change brings to winemaking is unpredictability. Producers used to know which varieties to grow, how to grow them, when to harvest the berries and how to ferment them to produce a consistent, quality wine – but today, every step is subject to uncertainty. In response to these challenges, winemakers are finding ways to preserve traditional as well as economically beneficial grape varieties and their unique qualities under the shifting conditions of global warming, and researchers are integrating knowledge, resources, and services regarding the grapevine (Navarro-Payá et al., 2022).
Temperature studies in vine are crucial as they help understand how variations in temperature affect vine development (de Rességuier et al., 2020; Merrill et al., 2020). Changes in temperature trigger shifts of ripening, which affect grape composition (Leeuwen and Darriet, 2016). For instance, higher temperatures have been found to correlate with flower abortion, potentially leading to decreased wine grape yields (Merrill et al., 2020). Another major challenge faced by grape growers is cold temperature damage, especially during the winter season. In many regions, grapevines are exposed to extreme cold, which can result in injury or death of the plants. Cold tolerance is a critical trait essential for the survival and productivity of grapevines in cold regions. However, the plant resistance to very cold and freezing temperatures has not been extensively studied yet despite recent advances in sequencing and molecular biotechnology (Ren et al., 2023). Also, not much is known about the impact of different temperatures on the gene expression patterns of Vitis vinifera (Han et al., 2023). Experiments done under various cold stress conditions showed that the freeze-shock damages plant leaves more than long-term freezing (Londo et al., 2018). The researchers found that the freeze-shock stress limits the sustainability and productivity of grapevines. The transcriptional landscape contrasts observed between low temperature and freezing stresses demonstrate quite different activation of candidate pathways impacting grapevine cold response. Genes from the ethylene signaling, abscisic acid signaling, the AP2, WRKY, and NAC transcription factor families, and starch/sucrose/galactose pathways were among the most observed to be differentially regulated. In response to cold stress, plants possess activation of specific metabolic pathways (including sugar accumulation and biosynthesis of prolines), changes in cell membrane rigidity, activation of calcium signaling and several ice-responding genes (like ICE1 and CBFs), and epigenetic regulation, to protect cells from ice nucleation, control cell membrane stability, scavenge reactive oxygen species, and adapt to cold stress, respectively (reviewed in Theocharis et al., 2012; Satyakam et al., 2022).
In Arabidopsis, the transcription factors AP2/ERF, WRKY, NAC, and MYB are known to enhance plant response to cold stress through various signaling pathways (reviewed in Abdullah et al., 2022). They are involved in the regulation of cold-responsive genes and contribute to increased cold tolerance. Specifically, MYB transcription factors are crucial in cold response due to their influence in regulation of cold-responsive genes, like CBF genes (Wang et al., 2023). Overexpression of certain MYB transcription factors can lead to enhanced tolerance to abiotic stress. Although the specific functions of these transcription factors in the cold acclimation of Arabidopsis are well-studied, there is only limited knowledge about their exact role in cold acclimation of grapevine.
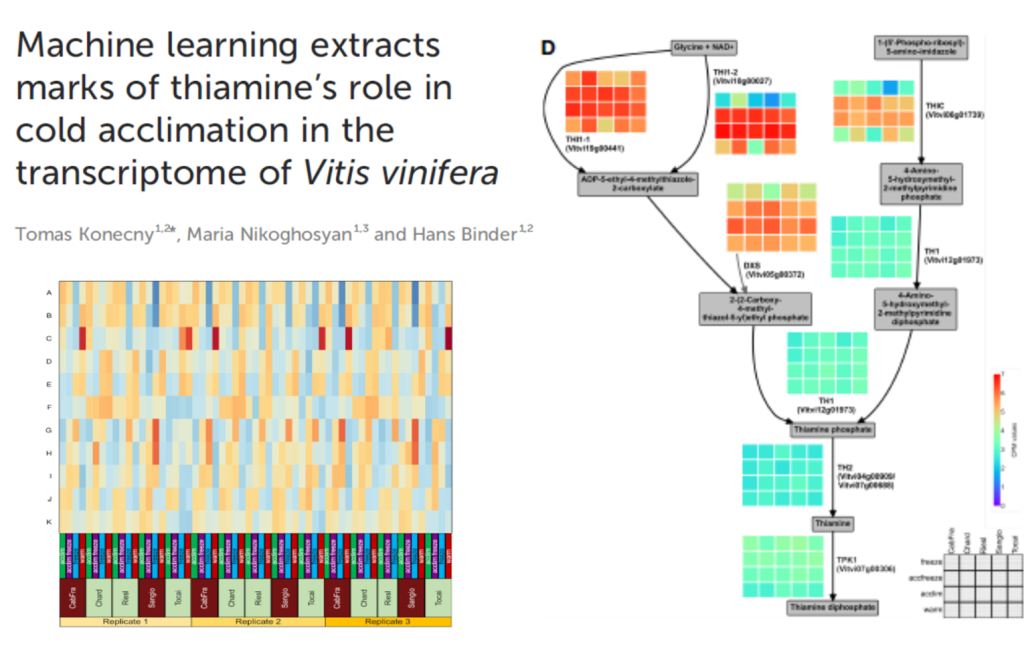
Epigenetic modifications, such as DNA methylation and post-translational modifications of histone proteins, play a crucial role in plant responses to cold stress (Sharma et al., 2022; Rehman et al., 2023). It is widely accepted that grapevine DNA methylation variability is primarily influenced by genotype. However, recent findings suggest that the environment where the grapevine grows can also significantly alter the methylome (Xie et al., 2017; Baránková et al., 2021). Epigenetic mechanisms are also associated with the regulation of metabolite biosynthesis and the accumulation of phenolic compounds in grapevines (He et al., 2010; Marfil et al., 2019). Thiamine, or vitamin B1, acts as a cofactor for several enzymes involved in metabolic pathways. It is crucial for plant health, particularly in defending against pathogens (Subki et al., 2018) and its role is known also in wine production (Bataillon et al., 1996). Thiamine has been found to boost the immunity and defense system of plants, playing a key role in their protection against biotic and abiotic stresses (Subki et al., 2018; Jaiwal et al., 2019). The exposure to abiotic stresses in the plant model organism Arabidopsis thaliana results in an upregulation of thiamin biosynthetic gene expression and the thiamine accumulation leading to enhanced tolerance to oxidative stress (Tunc-Ozdemir et al., 2009). However, the relationship between thiamine metabolism and temperature control in grapevines is not known. Our study aims to elucidate the expression of thiamin-biosynthetic genes in grapevines during cold acclimation and freezing stress.
Moreover, transcriptome-wide gene expression studies on larger sample sets are challenging because they aim at extracting relevant biological information such as affected pathways and related marker genes from a multidimensional data landscape with a co-variance structure much more complex than simple case-control settings. A series of machine-learning based methods such as weighted correlation network analysis (WGCNA) (Langfelder and Horvath, 2008) or non-negative matrix factorization (NMF) (Frigyesi and Höglund, 2008) have been developed to solve the problem via appropriate dimension reduction. Self-Organizing Maps (SOM) provide another option for knowledge mining in complex data to extract hidden covariance relations of reduced dimensionality.
The SOM machine learning method was developed by Kohonen over 30 years ago (Kohonen, 1982). It provides a very effective clustering algorithm which can be adapted to a wide range of applications (Loeffler-Wirth et al., 2020). We here make use of the “omics-portrayal” variant of SOM which combines supervised clustering of gene expression profiles into a two-dimensional grid of metagenes with unsupervised clustering into modules of co-regulated genes. These modules reflect the intrinsic co-variance landscape of the system in gene space. SOM portrayal possesses a series of advantages compared with alternative methods such as NMF or WCGNA (Wirth et al., 2011). Particularly, SOM-portrayal offers a comprehensive downstream analysis pipeline including different options for class discovery in sample and gene space, differential gene expression analysis, function and knowledge mining using gene set analysis with an implemented repository of more than five thousand gene signatures (Löffler-Wirth et al., 2015). SOM portrayal considers the multidimensional nature of gene regulation and pursues a modular view on co-expression, reduces dimensionality and, most importantly, supports visual perception in terms of individual, case-specific expression portraits. The pipeline has been applied to a series of data types and issues, e.g., in the context of molecular oncology (Binder et al., 2022; Loeffler-Wirth et al., 2022; Ashekyan et al., 2023) and health-related population studies (Nikoghosyan et al., 2019; Schmidt et al., 2020a), which all have proven the analytic strength of the methods in complex, multi-dimensional omics data. In the context of vine genomics, oposSOM has been applied so-far as “SOMmelier” to microarray SNP data to discover the dissemination history of Vitis vinifera as seen by vine genomes (Nikoghosyan et al., 2020). The main obstacle for applying the program to transcriptomic data of vine is the lack of gene annotations of their functional context. We here provide the first adaptation of oposSOM to vine transcriptomic data. We employed the SOM algorithm to uncover associations among the cold acclimation mechanisms that have not been characterized before.
In the context of grapevine species grown under different temperature conditions, SOM could help to identify differentially expressed genes of the thiamine biosynthetic pathway in response to temperature changes, providing valuable insights into molecular-level adaptations. Our analysis of transcriptomic mechanisms of temperature adaptation is in line with major questions that must be answered in the context of modern breeding practices nicknamed as Breeding 4.0 (Wallace et al., 2018), namely how do we adapt crops to better fit agricultural environments and what is the nature of the diversity upon which breeding can act?
. . .
RESULTS
Our analysis unveiled a connection with vitamin B1 (thiamine) biosynthesis, suggesting a link between temperature regulation and thiamine metabolism, in agreement with thiamine related stress response established in Arabidopsis before. Furthermore, we found that epigenetic mechanisms play a crucial role in regulating the expression of stress-responsive genes at low temperatures in grapevines.
to read the full article, click here: https://www.frontiersin.org/articles/10.3389/fpls.2023.1303542/full
